|
1/1/2024 0 Comments Calculate pi isoelectric point
Isoelectric point Proteomics pKa dissociation constant. This article was reviewed by Frank Eisenhaber and Zoltán Gáspári. So, to compute the isoelectric point: First, you add the pKa and pKb values. where: pKa Dissociation constant of acid pKb Dissociation constant of base and. The isoelectric point is of significance in protein purification because it is the pH at which solubility is often minimal and at which mobility in an electrofocusing system is zero, and therefore the point at which the protein. The pI can be estimated fairly accurately (within 0.1 or 0.2 pH units) from the pK values of all the proton dissociable groups in the peptide. The formula to calculate the isoelectric point of a substance is: pI (pKa + pKb) / 2. Below the isoelectric point proteins carry a net positive charge, above it a net negative charge. Draw the structures of the +1, 0, and -1 charged states of each amino acid. Step one: Consider the ionization of the amino acid. The isoelectric point (or pI) of an amino acid is the pH at which the amino acid has a net charge of zero. The IPC service is freely available at Peptide and protein datasets used in the study and the precalculated pI for the PDB and some of the most frequently used proteomes are available for large-scale analysis and future development. Isoelectric Point: The pH at which a protein carries no net charge. View the attachment below for the chemical structures. Moreover, the prediction of pI using the IPC pKa's leads to fewer outliers, i.e., predictions affected by errors greater than a given threshold. 
Each amino acid has a different dissociation constant, KD. The latter was determined using a protein isoelectric point. The isoelectric point is the pH at which the net charge is zero. 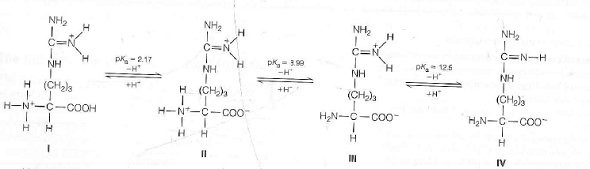
The isoelectric point is the pH at which the net. According to the presented benchmarks, the newly developed IPC pKa sets outperform previous algorithms by at least 14.9 % for proteins and 0.9 % for peptides (on average, 22.1 % and 59.6 %, respectively), which corresponds to an average error of the pI estimation equal to 0.87 and 0.25 pH units for proteins and peptides, respectively. calculate the predicted molecular weight and isoelectric (pI) point of all the proteins. An understanding of how to calculate isoelectric point is essential when you are working with proteins. Here, I present the Isoelectric Point Calculator (IPC), a web service and a standalone program for the accurate estimation of protein and peptide pI using different sets of dissociation constant (pKa) values, including two new computationally optimized pKa sets. For lysine and arginine, the isoelectric point, pI, occurs at a pH where the net charge on the nitrogen-containing groups is +1 and balances the charge of. Question: Calculate the pI (isoelectric point) for glutamate assuming that the pK1 is 2.19, the pKR is 4.25 and the pK2 is 9.67. Additionally, pI estimation can be helpful during protein crystallization trials. Calculate the pI (isoelectric point) for glutamate assuming that the pK1 is 2.19, the pKR is 4.25 and the pK2 is 9.67. With common personal computers and using a compiled version of the program, the time required to calculate pI values of macromolecules containing for example, 9. 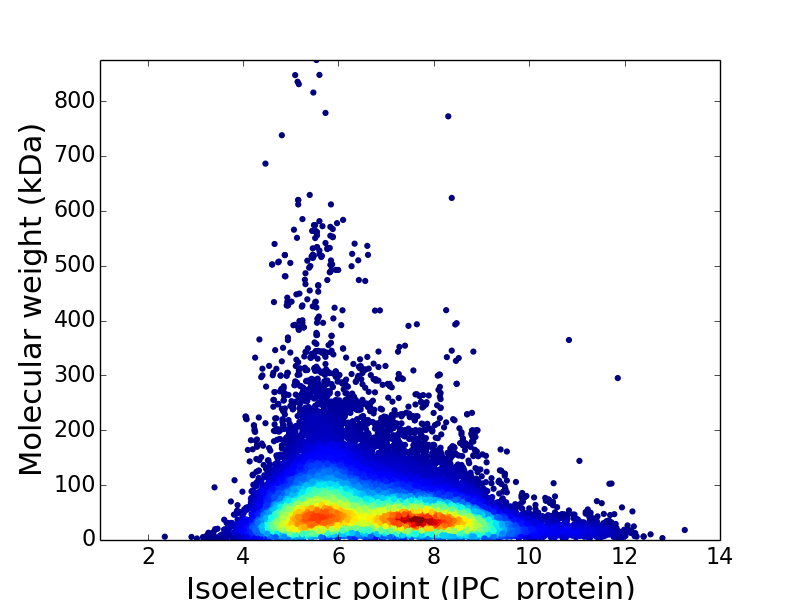
The $\mathrm pK_\mathrm a$ of the amino group is not relevant.Accurate estimation of the isoelectric point (pI) based on the amino acid sequence is useful for many analytical biochemistry and proteomics techniques such as 2-D polyacrylamide gel electrophoresis, or capillary isoelectric focusing used in combination with high-throughput mass spectrometry. Isoelectric point of an amino acid is the $\mathrm$ is simply the average of the two $\mathrm pK_\mathrm a$ values of the two carboxylic acid groups.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
